|
|
MEPSA: Minimum Energy Pathway Analysis for energy landscapes.
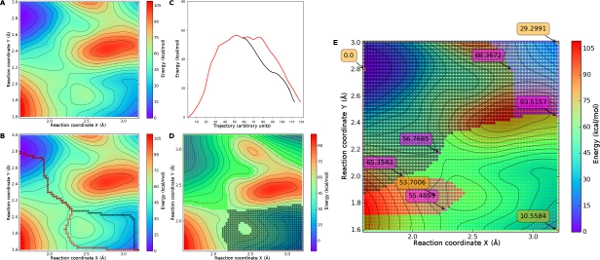
From conformational studies to atomistic descriptions of enzymatic reactions, potential and free energy landscapes can be used to describe biomolecular systems in detail. However, extracting the relevant data of complex 3D energy surfaces can sometimes be laborious. Here we present MEPSA (Minimum Energy Path Surface Analysis), a cross-platform user friendly tool for the analysis of energy landscapes from a transition state theory perspective. Some of its most relevant features are: identification of all the barriers and minima of the landscape at once, description of maxima edge profiles, detection of the lowest energy path connecting two minima and generation of transition state theory diagrams along these paths. In addition to a built-in plotting system, MEPSA can save most of the generated data into easily parseable text files, allowing more versatile uses of MEPSAs output such as the generation of molecular dynamics restraints from a calculated path.
Availability and implementation: MEPSA is freely available (under GPLv3 license) at this page: [http://bioweb.cbm.uam.es/software/MEPSA/]
CITATION: Marcos-Alcalde, I., Setoain, J., Mendieta-Moreno, J.I., Mendieta, J. & Gómez-Puertas, P. (2015). MEPSA: minimum energy pathway analysis for energy landscapes. Bioinformatics 31, 3853-3855. [https://doi.org/10.1093/bioinformatics/btv453].
PAPERS CITING MEPSA: [Google Scholar]
MEPSA exhibits a high citation profile: [Dimensions]
|
|
|
|
MEPSA DOWNLOAD (freely available, under GPLv3 license):
MEPSA v1.4 (Nov. 2019):
Perl script to convert a path into molecular dynamics restraints (Amber): [path_to_rst.zip]
DATA SAMPLES (MEPSA user manual):
ENERGY MAP -TEST- coordinates: [test_energy_map.zip]
Geneva-Turin MAP coordinates: [Geneva_Turin.zip]
New in version 1.4:
- Following a feature request from Dr. Robin A. Corey, if an error data column (a 4th column in the input file) is provided, it will be shown in energy profiles as well as in path saving txt files.
- Path saving txt files now indicate the index (counting from 0) of each point as it was read from the input file. Basically adding one to this index the user can obtain the line of the in input file from which each point of the resulting path/s comes.
FORMER VERSIONS (no longer supported):
MEPSA v1.3 (Nov. 2019)
MEPSA v1.3 source code (for Python 3.4.x): [mepsa_1.3_3.4x.zip]
MEPSA v1.3 user manual: [mepsa_manual_1.3.zip]
New in version 1.3:
- Deprecation fix: As reported by Dr. Robin A. Corey, griddata, which became deprecated in matplotlib, had to replaced by scipy.interpolate.griddata.
MEPSA v1.2 (Feb. 2018)
MEPSA v1.2 source code (for Python 3.4.x): [mepsa_1.2_3.4.x.zip]
MEPSA v1.2 user manual: [mepsa_manual_1.2.pdf]
New in version 1.2:
- Fixed well sampling for complex surfaces.
- Well sampling now only requires origin and target definition.
- Well sampling now only displays the wells sampled to reach the highest barrier connecting two states.
- New global connectivity analysis mode "MINIMAL": samples connectivity between states until every state is connected to another one.
- Bug-fix(minor): Modifying map with rectangle mode would print X and Y increments on terminal.
- Bug-fix(minor): saving well sampling to .txt files created files with wrong extension (.tx t instead of .txt).
MEPSA v1.1 (May, 2017)
MEPSA v1.1 source code (for Python 3.4.x): [mepsa_1.1_3.4.x.zip]
MEPSA v1.1 user manual: [mepsa_manual_1.1.pdf]
New in version 1.1:
- Many stability fixes regarding explicit int type specification in array indexes.
- During surface plots griddata now uses "linear" instead of "natural neighbour" interpolation, thus removing natgrid as a requirement.
- Adding points to plots is much faster now. Points are passed all at once in a 1D array instead of performing a slow for iterations.
- Python 2.7 no longer supported.
MEPSA v1.0 (Nov. 2015):
|
|
|
|
|
|
BIOWEB: MOLECULAR MODELLING GROUP
Centro de Biología Molecular "Severo Ochoa"
(CBMSO, CSIC-UAM)
http://bioweb.cbm.uam.es |
C/ Nicolás Cabrera, 1. Campus UAM, Cantoblanco
28049 Madrid. Spain
Tel: (+34) 91 196 4662 / 4663
Fax: (+34) 91 196 4420
|
|
|
|